How DNA Encoded Libraries Expedite Drug Discovery
Posted via | #Healthcare IT News: Drug Discovery #Healthcare IT News: Genomics
Drug discovery and development is a complex and tedious process requiring a significant amount of time and resources. In fact, on an average, the entire drug development process, from the demonstration of proof-of -concept to commercial launch, takes around 10-15 years and requires huge capital investments (USD 4-10 billion). Moreover, only a small fraction of early-stage therapeutic candidates are able to make it past preclinical testing, into clinical trials. Further, an even fewer number of clinical stage molecules are eventually approved for commercialization by the concerned regulatory authorities. Despite the advances in technology and improved understanding of biological systems, the drug discovery process is still considered to be lengthy, expensive, complex and inefficient.
Due to the aforementioned concerns, drug developers are gradually shifting their focus towards the use of novel discovery techniques, which include screening of combinatorial chemistry based large collections of compounds. Among all the screening techniques, DNA encoded library (DEL) technology has captured the attention of drug developers, owing to its various advantages, including rapid and effective screening of large number of compounds, reduced costs and time in drug discovery, low investment and less requirement of storage space and streamlined drug discovery process.
Overview of DNA Encoded Libraries
DNA encoded libraries have emerged as a robust and powerful tool, which combines the vast diversity of combinatorial chemistry and the versatile information encoding capabilities of DNA, to facilitate extremely high throughput molecular screening experiments. The use of DNA encoded libraries for hit generation is both cost effective and significantly expedites lengthy molecular screening processes. Fundamentally, the creation of a DNA encoded library is based on tagging small molecule / organic leads using short sequences of DNA and then using the combinatory chemistry approach to develop large numbers of structurally and functionally diverse molecules having unique DNA tags. Such a library can be synthesized within few weeks, typically containing millions to billions of compounds and using the following types of chemical reactions:
- Amide bond formation
- Nucleophilic aromatic substitutions
- Reductive amination
- Suzuki reactions
Screening experiments using such libraries are carried out in a single reaction tube for a singular biological target. In a typical screening experiment using a DEL, compounds that bind to the target are identified using their unique DNA tags and therefore easily isolated from the mixture. Such potential leads may be further incubated with the target, under varying conditions, in order to select the best candidate molecules, or hits. It is worth mentioning that the DNA tags on potential hits are designed to enable researchers to identify the reactions and molecular building blocks used for the synthesis of the same compounds.
Historical Evolution of DNA Encoded Libraries
The concept of a DNA encoded library was first proposed in 1992, by Richard Lerner (a chemist at the Scripps Research Institute California) and Sydney Brenner (cowinner of the 2002 Nobel Prize in Physiology or Medicine). Lerner and Brenner first published a paper based on thought experiments, highlighting the advantages of an encoded molecular library. Later, the scientists developed a small DNA encoded library and patented it under US patent no. 5573905. Although the proposition was widely acknowledged by the research community, the idea remained mostly dormant for about a decade, till the necessary technology and information base was developed to support the application of this revolutionary concept. Nuevolution was one of the first companies to initiate the development of DNA encoded libraries, other pioneering players in this field include HitGen and X-Chem Pharmaceuticals.
Figure below presents a historical timeline highlighting the evolution of DNA encoded libraries.

DELs vs. Traditional Screening methods in Drug Discovery
As mentioned earlier, the use of DNA encoded libraries enables the development and optimization of new drug compounds as well as it also aids in the generation of valuable data related to their structure and interactions with different biological molecules.
Figure below provides the difference between traditional libraries and DNA encoded libraries.
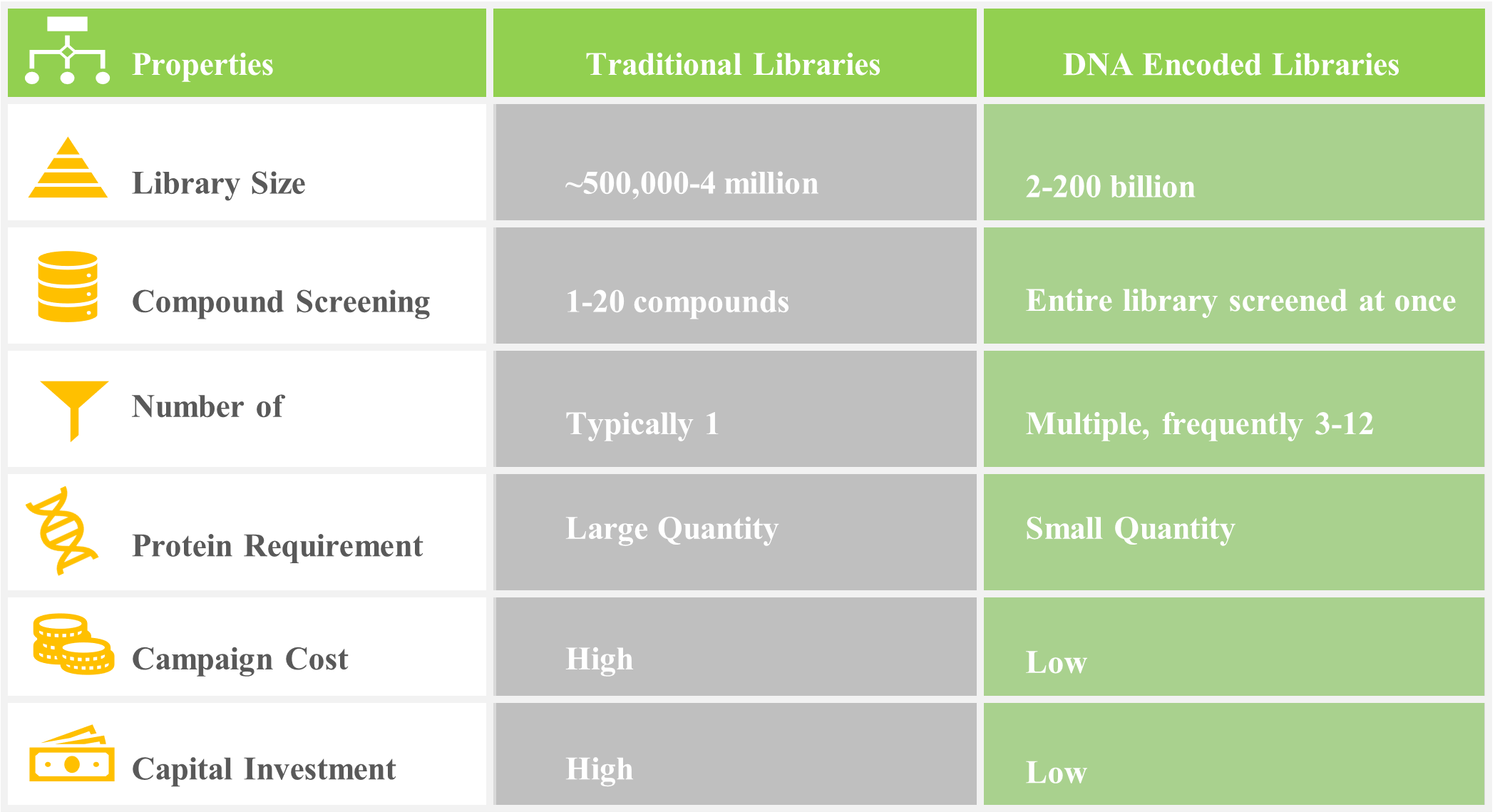
Benefits of using DELs in Complex Drug Development
The use of DNA encoded library technology is very helpful during the early stages of drug discovery since it requires less investment, time and storage space to identify target compounds. Apart from these, there are several other advantages as well associated with the application of DNA encoded libraries in accelerating drug development.
Figure below presents a summary DEL technology benefits.
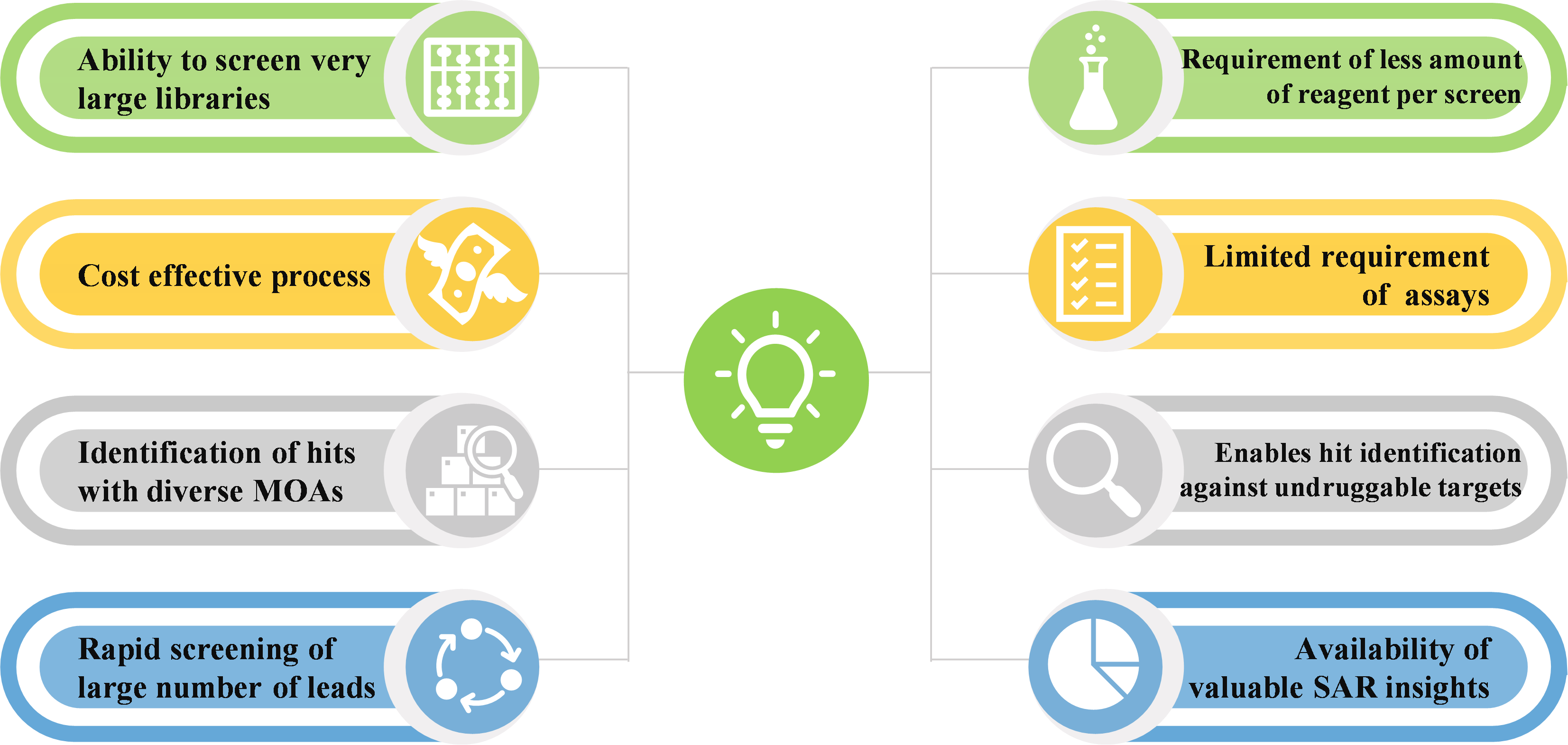
Challenges and Limitations Associated with DNA Encoded Libraries
Despite having numerous advantages, DNA encoded libraries are fraught with various challenges. Figure below presents a summary of the challenges / limitations associated with DNA encoded libraries.
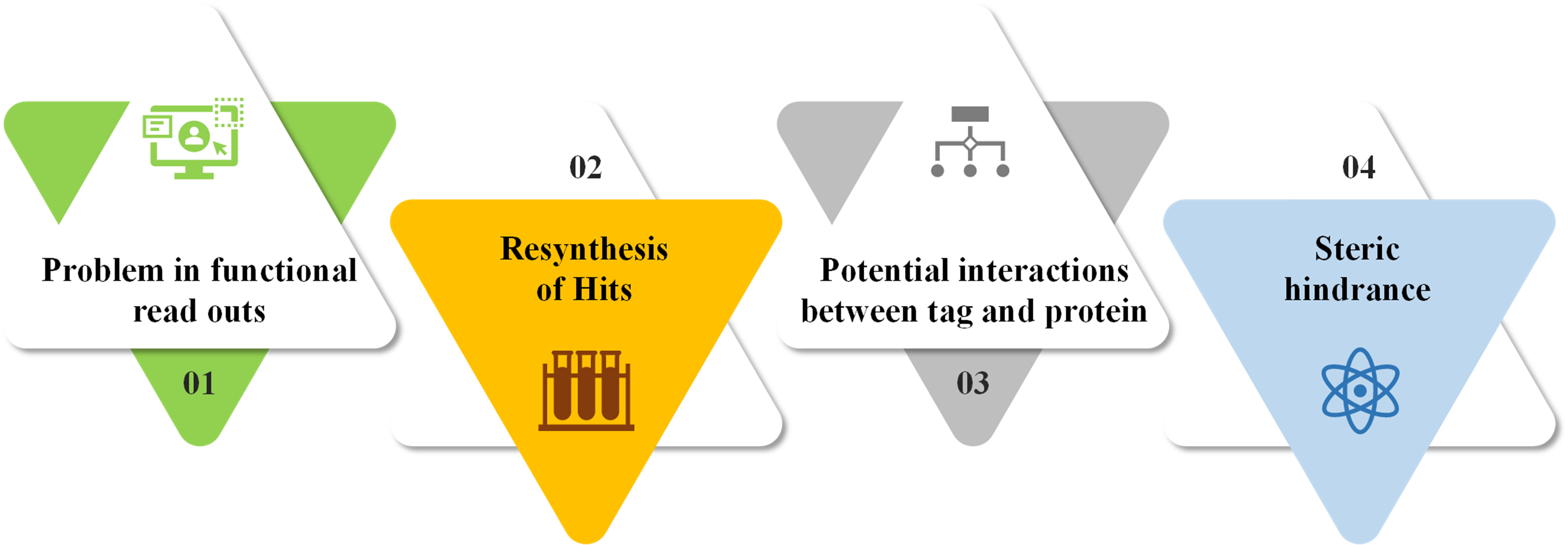
Future Perspectives and Opportunity Areas
The identification of suitable molecules that are capable of binding to specific biological targets and mediating a therapeutic effect is the primary challenge in the field of chemistry, biology and medicine. As a result, many research groups are engaged in exploring technologies that can efficiently facilitate the identification of ligands for drug discovery. DNA encoded libraries represent one of the latest and most versatile methods of molecular screening, which enables high throughput screening of large libraries of compounds in an inexpensive and timebound manner. Functioning analogous to the phage display technique, which is used for antibody development, this method of hit-generation and screening has revolutionized the small molecule drug discovery.
Over time, a variety of DNA encoded library construction strategies have been developed and investigated by both academic and industry players. With the aid of revolutionary artificial intelligence (AI) and machine learning (ML) tools, the process of DNA encoded libraries is being further streamlined using algorithms to navigate through these large datasets of compounds and predict the best drug targets. Although DNA encoded libraries have been successful in enabling the identification and isolation of hits against several types of target proteins, certain studies have reported their inability to be used to interrogate certain protein classes. Therefore, even though these libraries have gradually been accepted into mainstream drug discovery applications, there is still scope for further improvement.
Experts believe that once the abovementioned challenges are addressed, it is likely to open up new opportunities for DNA encoded libraries. It is worth mentioning that the scientific community is actively petitioning for this technology to be made open source. It is, therefore, apt to assume that the future may witness greater interactions between public and private research groups and library developers / providers, focused on the discovery of pharmacological leads against a wide array of biological targets.
Continue reading at | #Healthcare IT News: Drug Discovery #Healthcare IT News: Genomics
See Also
- Health Care Job Market Expands in August
- Oncology Software: The Digital Backbone of Cancer Care
- Why It Is Important To Understand Multimodal Large Language Models In Healthcare
- A Blueprint Is Essential To Accelerate Digital Transformation And Improve Patient Outcomes
- The Medical Library: A Hospital’s Most Underappreciated Asset